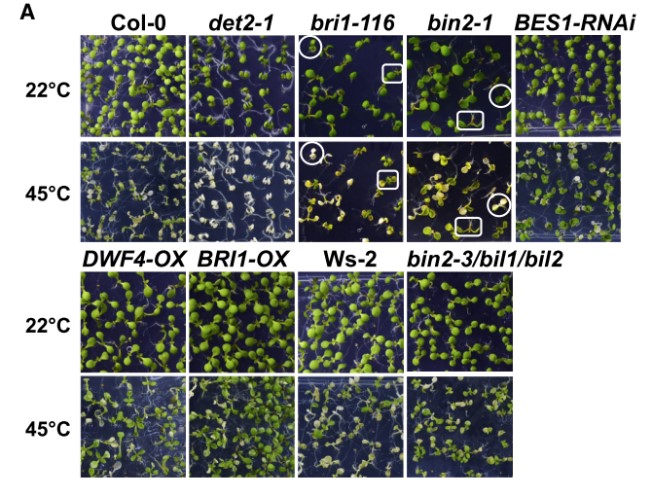
Brassinosteroids promote thermotolerance through releasing BIN2-mediated phosphorylation and suppression of HsfA1 transcription factors in Arabidopsis
High temperature adversely affects plant growth and development. The steroid phytohormones brassinos- teroids (BRs) are recognized to play important roles in plant heat stress responses and thermotolerance, but the underlying mechanisms remain obscure. Here, we demonstrate that the glycogen synthase kinase 3 (GSK3)-like kinase BRASSINOSTEROID INSENSITIVE2 (BIN2), a negative component in the BR signaling pathway, interacts with the master heat-responsive transcription factors CLASS A1 HEAT SHOCK TRAN- SCRIPTION FACTORS (HsfA1s). Furthermore, BIN2 phosphorylates HsfA1d on T263 and S56 to suppress its nuclear localization and inhibit its DNA-binding ability, respectively. BR signaling promotes plant ther- motolerance by releasing the BIN2 suppression of HsfA1d to facilitate its nuclear localization and DNA binding. Our study provides insights into the molecular mechanisms by which BRs promote plant thermo- tolerance by strongly regulating HsfA1d through BIN2 and suggests potential ways to improve crop yield under extreme high temperatures.(Plant Communications)
Variation in cis-regulation of a NAC transcription factor contributes to drought tolerance in wheat
Drought is a major environmental factor limiting wheat production worldwide, and developing drought- tolerant cultivars is a central challenge for wheat breeders globally. Therefore, it is important to identify genetic components determining drought tolerance in wheat. In this study, we identified a wheat NAC gene (TaNAC071-A) that is tightly associated with drought tolerance by a genome-wide association study. Knockdown of TaNAC071-A in wheat attenuated plant drought tolerance, whereas its overexpression significantly enhanced drought tolerance through improved water-use efficiency and increased expression of stress-responsive genes. This heightened water-saving mechanism mitigated the yield loss caused by water deficit. Further candidate gene association analysis showed that a 108-bp insertion in the promoter of TaNAC071-A alters its expression level and contributes to variation in drought tolerance among wheat accessions. This insertion contains two MYB cis-regulatory elements (CREs) that can be directly bound by the MYB transcription activator, TaMYBL1, thereby leading to increased TaNAC071-A expression and plant drought tolerance. Importantly, introgression of this 108-bp insertion allele, TaNAC071-AIn-693, into drought-sensitive cultivars could improve their drought tolerance, demonstrating that it is a valuable genetic resource for wheat breeding. Taken together, our findings highlight a major breakthrough in determining the genetic basis underlying phenotypic variation in wheat drought tolerance and showcase the potential of exploiting CRE-containing indels for improving important agronomical traits.(Molecular Plant)

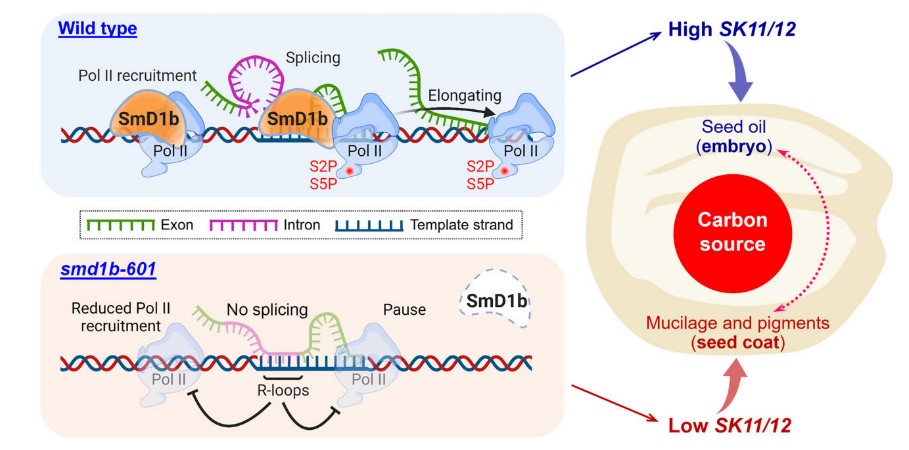
Splicing-mediated activation of SHAGGY-like kinases underpinning carbon partitioning in Arabidopsis seeds
Glycogen synthase kinase 3 (GSK3) family members serve as signaling hubs for plant development and stress responses, yet the underlying mechanism of their transcriptional regulation remains a long-standing mystery. Here we show that the tran- scription of SHAGGY-like kinase 11/12 (SK11/12), two members of the GSK3 gene family, is promoted by the splicing factor SmD1b, which is essential for distributing carbon sources into storage and protective components in Arabidopsis seeds. The chromatin recruitment of SmD1b at the SK11/12 loci promotes their transcription associated with co-transcriptional splicing of the first introns in the 50-untranslated region of SK11/12. The loss of SmD1b function generates transcripts with unspliced introns that create disruptive R-loops to hamper the transcriptional elongation of SK11/12, in addition to compromising the recruitment of RNA polymerase II to the SK11/12 genomic regions. These effects imposed by SmD1b de- termine the transcription of SK11/12 to confer a key switch of carbon flow among metabolic pathways in zygotic and ma- ternal tissues in seeds.(The plant cell)
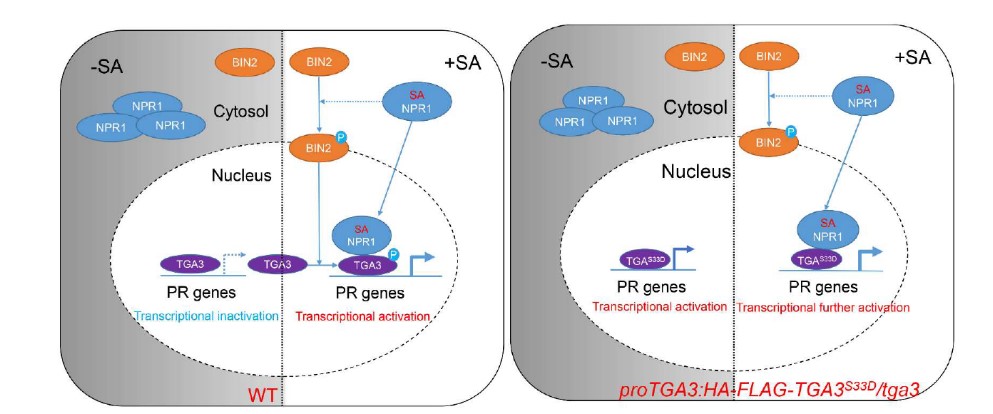
Salicylic acid-activated BIN2 phosphorylation of TGA3 promotes Arabidopsis PR gene expression and disease resistance
The plant defense hormone, salicylic acid (SA), plays essential roles in immunity and systemic acquired resistance. Salicylic acid induced by the pathogen is perceived by the receptor nonexpressor of pathogenesis-related genes 1 (NPR1), which is recruited by TGA transcription factors to induce the expression of pathogenesis- related (PR) genes. However, the mechanism by which post- translational modifications affect TGA’s transcriptional activity by salicylic acid signaling/pathogen infection is not well-established. Here, we report that the loss-of-function mutant of brassinos- teroid insensitive2 (BIN2) and its homologs, bin2-3 bil1 bil2, causes impaired pathogen resistance and insensitivity to SA-induced PR gene expression, whereas the gain-of-function mutant, bin2-1, exhibited enhanced SA signaling and immunity against the pathogen. Our results demonstrate that salicylic acid activates BIN2 kinase, which in turn phosphorylates TGA3 at Ser33 to enhance TGA3 DNA binding ability and NPR1–TGA3 complex forma- tion, leading to the activation of PR gene expression. These find- ings implicate BIN2 as a new component of salicylic acid signaling, functioning as a key node in balancing brassinosteroid-mediated plant growth and SA-induced immunity. (The EMBO Journal)
Insights into breeding history, hotspot regions of selection and untapped allelic diversity for bread wheat breeding
Breeding has increasingly altered the genetics of crop plants since the domestication of their wild progenitors. It is postulated that the genetic diversity of elite wheat breeding pools is too narrow to cope with future challenges. In contrast, plant genetic resources (PGRs) of wheat stored in gene banks are valuable sources of unexploited genetic diversity. Therefore, to ensure breeding progress in the future, it is of prime importance to identify the useful allelic diversity available in PGRs and to transfer it into elite breeding pools. Here, a diverse collection consisting of modern winter wheat cultivars and gene bank accessions was investigated based on reduced-representation genomic sequencing and an iSelect single nucleotide polymorphism (SNP) chip array. Analyses of these datasets provided detailed insights into population structure, levels of genetic diversity, sources of new allelic diversity and genomic regions affected by breeding activities. We identified 57 regions representing genomic signatures of selection and 827 regions representing private alleles associated exclusively with genebank accessions. The presence of known functional wheat genes, quantitative trait loci and large chromosomal modifications, i.e., introgressions from wheat wild relatives, provided initial evidence for putative traits associated within these identified regions. These findings were supported by the results of ontology enrichment analyses. The results reported here will stimulate further research, and promote breeding in the future by allowing for the targeted introduction of novel allelic diversity into elite wheat breeding pools.(The Plant Journal)
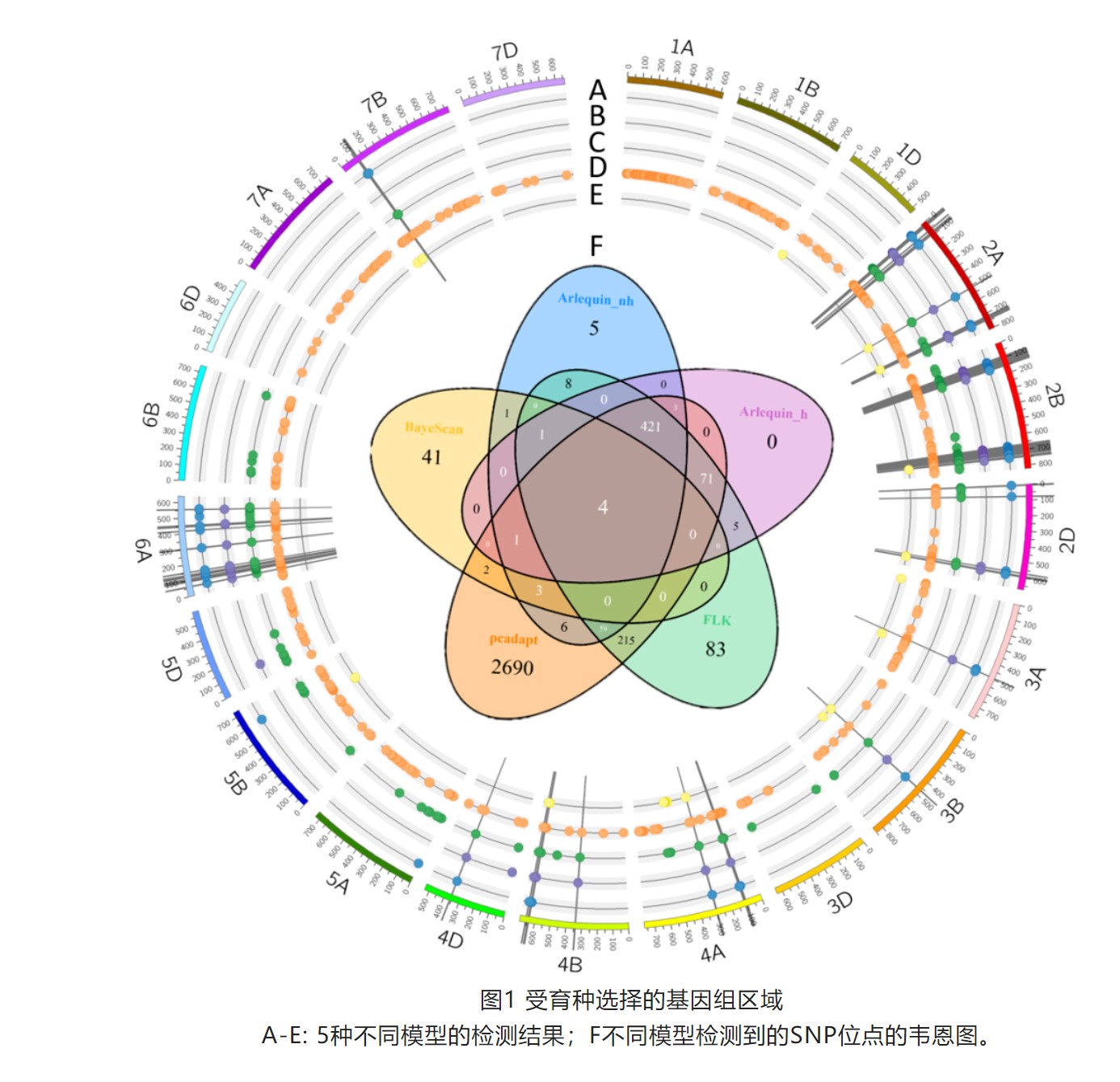
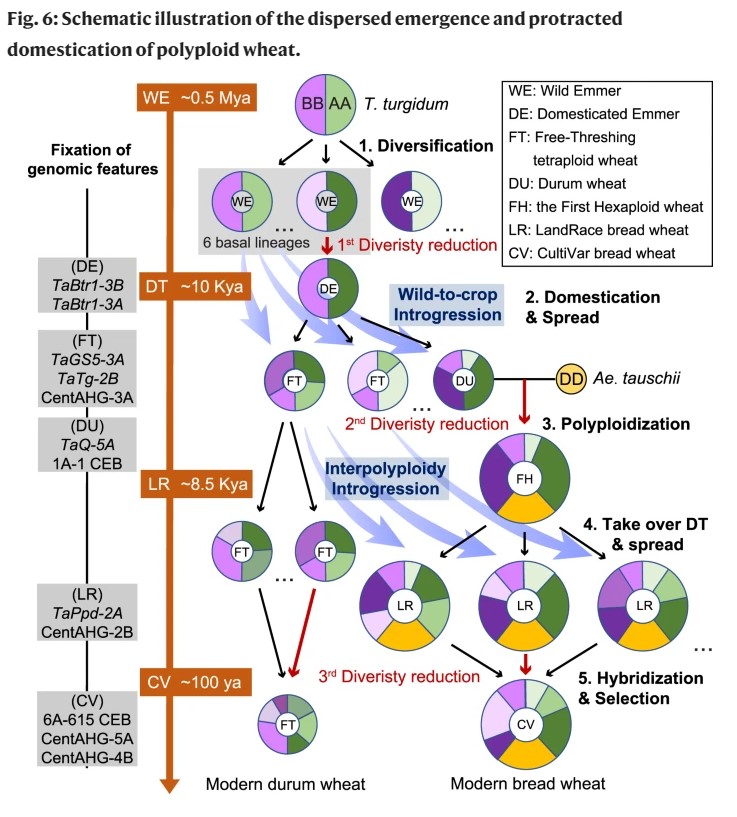
Whole-genome analysis of hard winter wheat germplasm identifies genomic regions associated with spike and kernel traits
Genetic dissection of yield component traits including spike and kernel characteristics is essential for the continuous improvement in wheat yield. Genome-wide association studies (GWAS) have been frequently used to identify genetic determinants for spike and kernel-related traits in wheat, though none have been employed in hard winter wheat (HWW) which represents a major class in US wheat acreage. Further, most of these studies relied on assembled diversity panels instead of adapted breeding lines, limiting the transferability of results to practical wheat breeding. Here we assembled a population of advanced/elite breeding lines and well-adapted cultivars and evaluated over four environments for phenotypic analysis of spike and kernel traits. GWAS identified 17 significant multi-environment marker–trait associations (MTAs) for various traits, representing 12 putative quantitative trait loci (QTLs), with five QTLs affecting multiple traits. Four of these QTLs mapped on three chromosomes 1A, 5B, and 7A for spike length, number of spikelets per spike (NSPS), and kernel length are likely novel. Further, a highly significant QTL was detected on chromosome 7AS that has not been previously associated with NSPS and putative candidate genes were identified in this region. The allelic frequencies of important quantitative trait nucleotides (QTNs) were deduced in a larger set of 1,124 accessions which revealed the importance of identified MTAs in the US HWW breeding programs. The results from this study could be directly used by the breeders to select the lines with favorable alleles for making crosses, and reported markers will facilitate marker-assisted selection of stable QTLs for yield components in wheat breeding.(Theor Appl Genet)
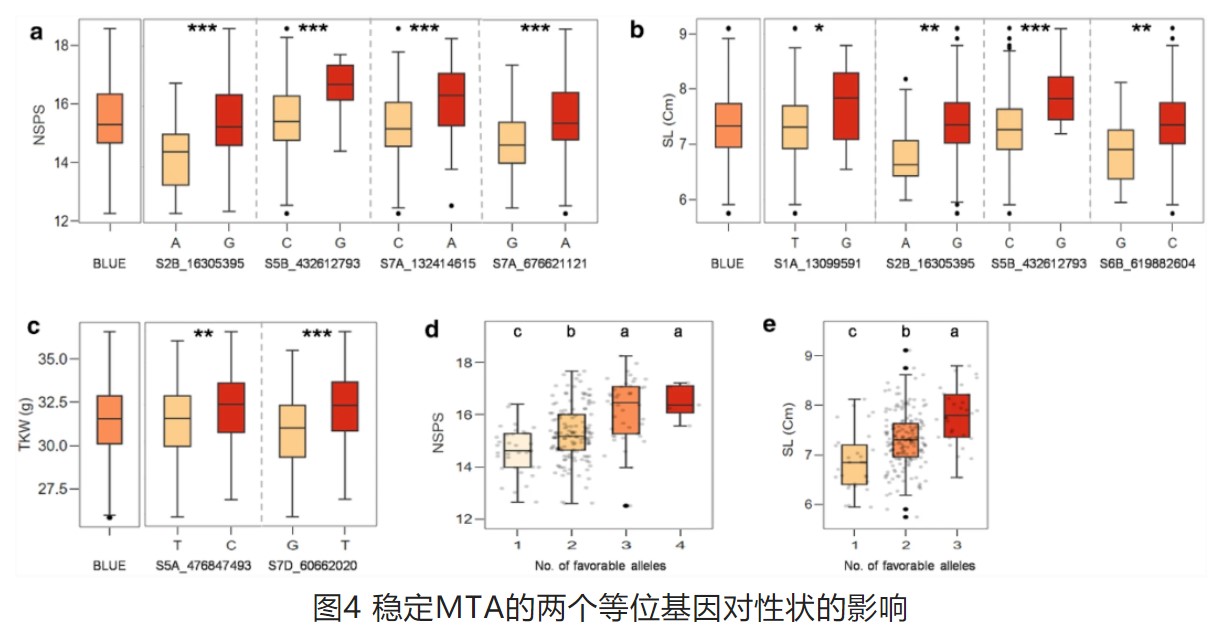
Dispersed emergence and protracted domestication of polyploid wheat uncovered by mosaic ancestral haploblock inference
Major crops are all survivors of domestication bottlenecks. Studies have focused on the genetic loci related to the domestication syndrome, while the contribution of ancient haplotypes remains largely unknown. Here, an ancestral genomic haploblock dissection method is developed and applied to a resequencing dataset of 386 tetraploid/hexaploid wheat accessions, generating a pan-ancestry haploblock map. Together with cytoplastic evidences, we reveal that domesticated polyploid wheat emerged from the admixture of six founder wild emmer lineages, which contributed the foundation of ancestral mosaics. The key domestication-related loci, originated over a wide geographical range, were gradually pyramided through a protracted process. Diverse stable-inheritance ancestral haplotype groups of the chromosome central zone are identified, revealing the expanding routes of wheat and the trends of modern wheat breeding. Finally, an evolution model of polyploid wheat is proposed, highlighting the key role of wild-to-crop and interploidy introgression, that increased genomic diversity following bottlenecks introduced by domestication and polyploidization.( nature communications)
Genome architecture and diverged selection shaping pattern of genomic differentiation in wild barley(Triticum aestivum)
Divergent selection of populations in contrasting environments leads to functional genomic divergence. However, the genomic architecture underlying heterogeneous genomic differentiation remains poorly understood. Here, we de novo assembled two high-quality wild barley (Hordeum spontaneum K. Koch) genomes and examined genomic differentiation and gene expression patterns under abiotic stress in two populations. These two populations had a shared ancestry and originated in close geographic proximity but experienced different selective pressures due to their contrasting micro-environments. We identified structural variants that may have played significant roles in affecting genes potentially associated with well-differentiated phenotypes such as flowering time and drought response between two wild barley genomes. Among them, a 29-bp insertion into the promoter region formed a cis-regulatory element in the HvWRKY45 gene, which may contribute to enhanced tolerance to drought. A single SNP mutation in the promoter region may influence HvCO5 expression and be putatively linked to local flowering time adaptation. We also revealed significant genomic differentiation between the two populations with ongoing gene flow. Our results indicate that SNPs and small SVs link to genetic differentiation at the gene level through local adaptation and are maintained through divergent selection. In contrast, large chromosome inversions may have shaped the heterogeneous pattern of genomic differentiation along the chromosomes by suppressing chromosome recombination and gene flow. Our research offers novel insights into the genomic basis underlying local adaptation and provides valuable resources for the genetic improvement of cultivated barley.(Plant Biotechnol J)
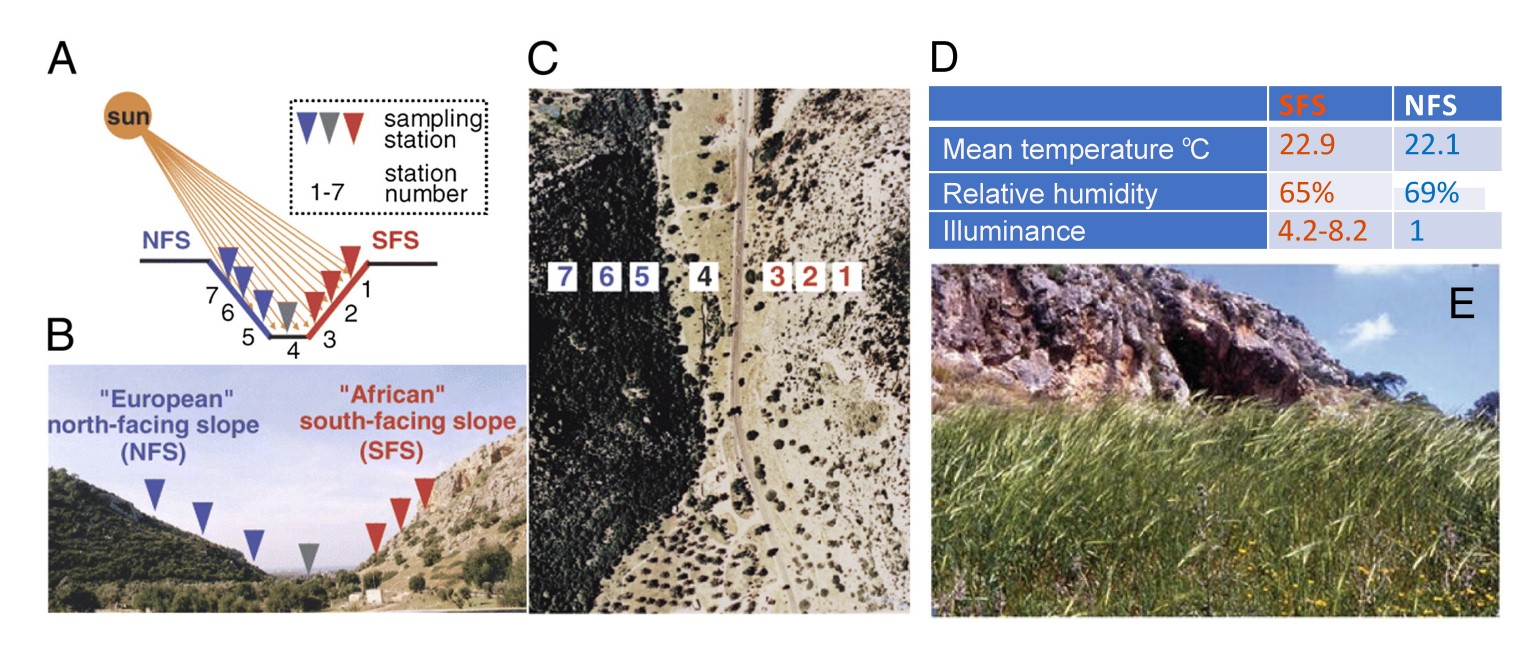

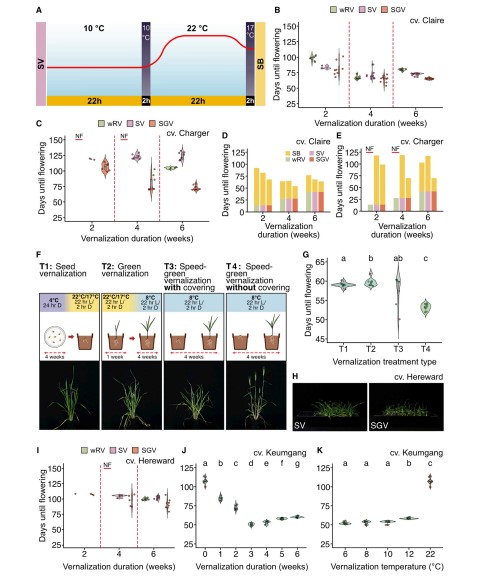
Speed vernalization to accelerate generation advance in winter cereal crops
There are many challenges facing the development of high-yielding, nutritious crops for future environments. One limiting factor is generation time, which prolongs research and plant breeding timelines. Recent advances in speed breeding protocols have dramatically reduced generation time for many short-day and long-day species by optimizing light and temperature conditions during plant growth. However, winter crops with a vernalization requirement still require up to 6–10 weeks in low-temperature conditions before the transition to reproductive development. Here, we tested a suite of environmental conditions and protocols to investigate whether the vernalization process can be accelerated. We identified a vernalization method consisting of exposing seeds at the soil surface to an extended photoperiod of 22 h day:2 h night at 10C with transfer to speed breeding conditions that dramatically reduces generation time in both winter wheat (Triticum aestivum) and winter barley (Hordeum vulgare). Implementation of the speed vernalization protocol followed by speed breeding allowed the completion of up to five generations per year for winter wheat or barley, whereas only two generations can be typically completed under standard vernalization and plant growth conditions. The speed vernalization protocol developed in this study has great potential to accelerate biological research and breeding outcomes for winter crops.(Molecular Plant)
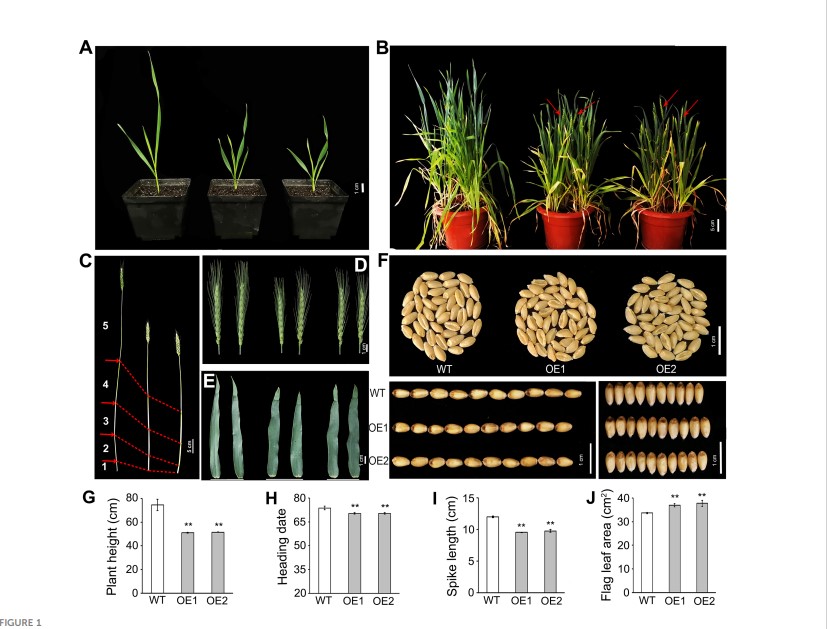
Overexpression of TaLBD16-4D alters plant architecture and heading date in transgenic wheat
Lateral organ boundaries domain (LBD) proteins, a class of plant-specific transcription factors with a special domain of lateral organ boundaries (LOB), play essential roles in plant growth and development. However, there is little known about the functions of these genes in wheat to date. Our previous study demonstrated that TaLBD16-4D is conducive to increasing lateral root number in wheat. In the present work, we further examined important agronomical traits of the aerial part of transgenic wheat overexpressing TaLBD16-4D. Interestingly, it was revealed that overexpressing TaLBD16-4D could lead to early heading and multiple alterations of plant architecture, including decreased plant height, increased flag leaf size and stem diameter, reduced spike length and tillering number, improved spike density and grain width, and decreased grain length. Moreover, auxin-responsive experiments demonstrated that the expression of TaLBD16-4D in wild-type (WT) wheat plants showed a significant upregulation through 2,4-D treatment. TaLBD16- 4D-overexpression lines displayed a hyposensitivity to 2,4-D treatment and reduced shoot gravitropic response. The expressions of a set of auxinresponsive genes were markedly different between WT and transgenic plants. In addition, overexpressing TaLBD16-4D affected the transcript levels of flowering-related genes (TaGI, TaCO1, TaHd1, TaVRN1, TaVRN2, and TaFT1). Notably, the expression of TaGI, TaCO1, TaHd1, TaVRN1, and TaFT1 displayed significant upregulation under IAA treatment. Collectively, our observations indicated that overexpressing TaLBD16-4D could affect aerial architecture and heading time possibly though participating in the auxin pathway.(Plant Science )
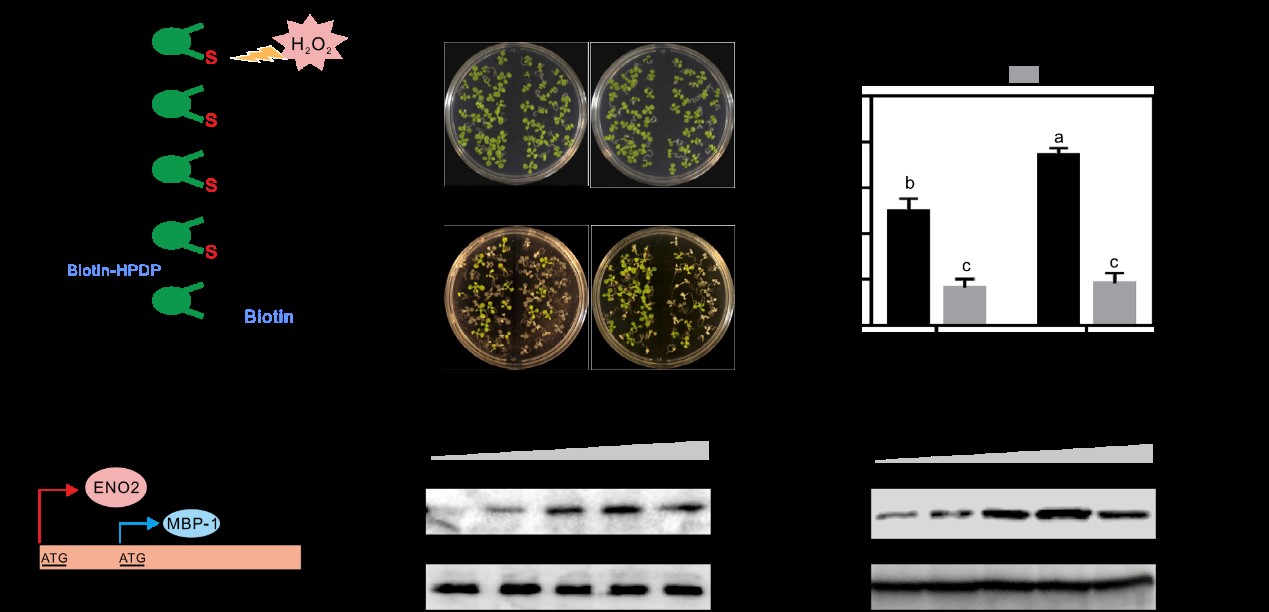
Sulfenylation of ENOLASE2 facilitates H2O2-conferredfreezing tolerance in Arabidopsis
H2O2 affects the expression of genes that are involved in plant responses to diverse environmental stresses;however, the underlying mechanisms remain elusive. Here, we demonstrate that H2O2 enhances plant freezing tolerance through its effect on a protein product of low expression of osmotically responsive genes2 (LOS2).LOS2 is translated into a major product, cytosolic enolase2 (ENO2), and sometimes an alternative product, the transcription repressor c-Myc-binding protein (MBP-1). ENO2, but not MBP-1, promotes cold tolerance by binding the promoter of C-repeat/DRE binding factor1 (CBF1), a central transcription factor in plant cold signaling, thus activating its expression. Overexpression of CBF1 restores freezing sensitivity of a LOS2loss-of-function mutant. Furthermore, cold-induced H2O2 increases nuclear import and transcriptional binding activity of ENO2 by sulfenylating cysteine 408 and thereby promotes its oligomerization. Collectively, our results illustrate how H2O2 activates plant cold responses by sulfenylating ENO2 and promoting its oligomerization, leading to enhanced nuclear translocation and transcriptional activation of CBF1.(Developmental Cell)
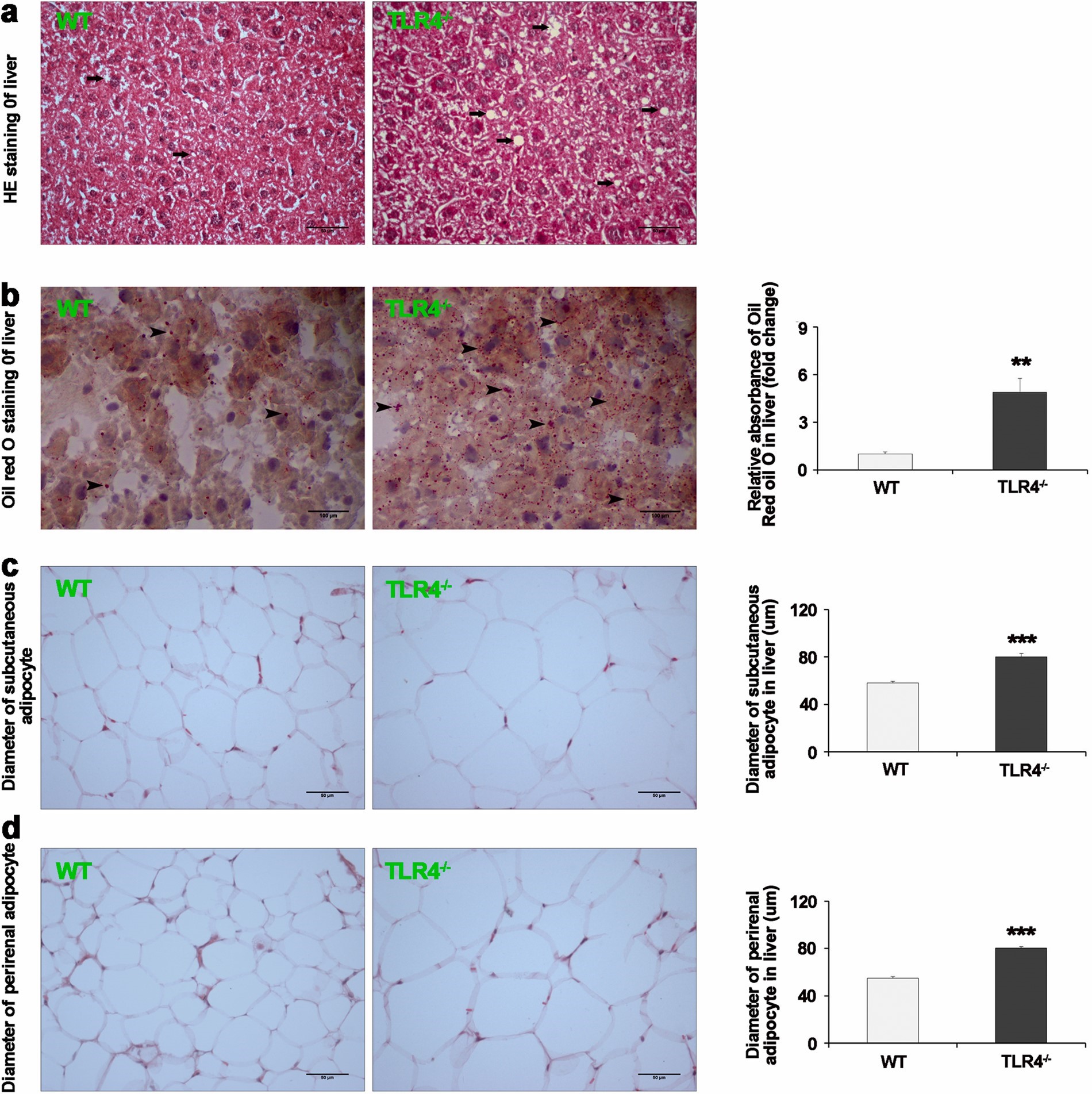
Chronic low-grade inflammation is involved in TLR4 knockout-induced spontaneous obesity in aged mice
Chronic inflammation plays an important role in obesity-related complications, including insulin resistance, type 2 diabetes, and cardiovascular disease. The imbalances between T helper (Th)1/Th2 cells and Th17/regulatory T (Treg) cells participate in the pathogenesis of inflammation. Previously it was demonstrated that Toll-like receptor (TLR) 4 knockout (KO) prevents high-fat diet (HFD)-induced obesity of young mice (6 months of age), however the effect of TLR4 KO on spontaneous obesity in aged mice (18 month of age) is still unknown. To further study this, TLR4 KO and WT mice were fed with a standard chow diet from weaning to the endpoint of the experiment. We found that TLR4-/- mice were thinner compared with WT mice at 6 months (M) old. However, TLR4-/- mice spontaneously developed obesity with increased weight and adiposity in both subcutaneous and visceral fat depots by 18 M old. Our results also indicated that TLR4 KO activated TRIF/IRF3 signalling, induced inflammation, and repolarised alternatively-activated (M2) macrophages to classically-activated (M1) macrophages. In addition, TLR4 KO resulted in an increased spleen index and induced imbalances of Th1/Th2 and Th17/Treg cells which indicated the occurrence of chronic low-grade inflammation. In conclusion, chronic low- grade inflammation induced by TLR4 KO was involved in spontaneous obesity in aged mice. An emerging link was established among the TRIF/IRF3 pathway, chronic low-grade inflammation, and obesity. We hope that these novel findings will provide a potential preventive strategy for obesity and build a spontaneous obesity mouse model.(Biomedicine & Pharmacotherapy)
Glutathione S-Transferase Interactions Enhance Wheat Resistance to Powdery Mildew but Not Wheat Stripe
Wheat stripe rust and powdery mildew are important worldwide diseases of wheat (Triticum aestivum). The wheat cultivar Xingmin318 (XM318) is resistant to both wheat stripe rust and powdery mildew, which are caused by Puccinia striiformis f. sp. tritici (Pst) and Blumeria graminis f. sp. tritici (Bgt), respectively. To explore the difference between wheat defense response against Pst and Bgt, quantitative proteomic analyses of XM318 inoculated with either Pst or Bgt were performed using tandem mass tags (TMT) technology. A total of 741 proteins were identified as differentially accumulated proteins (DAPs). Bioinformatics analyses indicated that some functional categories, including antioxidant activity and immune system process, exhibited obvious differences between Pst and Bgt infections. Intriguingly, only 42 DAPs responded to both Pst and Bgt infections. Twelve DAPs were randomly selected for RT-qPCR analysis, and the mRNA expression levels of 11 were consistent with their protein expression. Furthermore, gene silencing using the virus-induced gene silencing system indicated that glutathione S-transferase (TaGSTU6) has an important role in resistance to Bgt but not to Pst. TaGSTU6 interacted with the cystathionine beta-synthase (CBS) domain-containing protein (TaCBSX3) in both Pst and Bgt infections. Knockdown of TaCBSX3 expression only reduced wheat resistance to Bgt infection. Overexpression of TaGSTU6 and TaCBSX3 in Arabidopsis (Arabidopsis thaliana) promoted plant resistance to Pseudomonas syringae pv. Tomato DC3000. Our results indicate that TaGSTU6 interaction with TaCBSX3 only confers wheat resistance to Bgt, suggesting that wheat has different response mechanisms to Pst and Bgt stress.(Plant Physiology)
PIF7 is a master regulator of thermomorphogenesis in shade
Exploring the genetic variability in yield and yield-related traits is essential to continue improving genetic gains. Fifty-nine Argentinian durum wheat cultivars were analyzed for important agronomic traits in three field experiments. The collection was genotyped with 3565 genome-wide SNPs and functional markers in order to determine the allelic variation at Rht-B1 and Ppd-A1 genes. Population structure analyses revealed the presence of three main groups, composed by old, modern and genotypes with European or CIMMYT ancestry. The photoperiod sensitivity Ppd-A1b allele showed higher frequency (75%) than the insensitivity one Ppd-A1a (GS105). The semi-dwarfism Rht-B1b and the Ppd-A1a (GS105) alleles were associated with increases in harvest index and decreases in plant height, grain protein content and earlier heading date, although only the varieties carrying the Rht-B1 variants showed differences in grain yield. Out of the two main yield components, grain number per plant was affected by allelic variants at Rht-B1 and Ppd-A1 loci, while no differences were observed in thousand kernel weight. The increases in grain number per spike associated with Rht-B1b were attributed to a higher grain number per spikelet, whereas Ppd-A1a (GS105) was associated with higher grain number per spikelet, but also with lower spikelets per spike.(Nature Commun)
Root angle is controlled by EGT1 in cereal crops employing an antigravitropic mechanism
Root angle in crops represents a key trait for efficient capture of soil resources. Root angle is determined by competing gravitropic versus antigravitropic offset (AGO) mechanisms. Here we report a root angle regulatory gene termed ENHANCED GRAVI-TROPISM1 (EGT1) that encodes a putative AGO component, whose loss-of-function enhances root gravitropism. Mutations in barley and wheat EGT1 genes confer a striking root phenotype, where every root class adopts a steeper growth angle. EGT1 encodes an F-box and Tubby domain-containing protein that is highly conserved across plant species. Haplotype analysis found that natural allelic variation at the barley EGT1 locus impacts root angle. Gravitropic assays indicated that Hvegt1 roots bend more rapidly than wild-type. Transcript profiling revealed Hvegt1 roots deregulate reactive oxygen species (ROS) homeostasis and cell wall-loosening enzymes and cofactors. ROS imaging shows that Hvegt1 root basal meristem and elongation zone tissues havereduced levels. Atomic force microscopy measurements detected elongating Hvegt1 root cortical cell walls are significantly less stiff than wild-type. In situ analysis identified HvEGT1 is expressed in elongating cortical and stele tissues, which are distinct from known root gravitropic perception and response tissues in the columella and epidermis,respectively. We propose that EGT1 controls root angle by regulating cell wall stiffnessin elongating root cortical tissue, counteracting the gravitropic machinery’s known ability to bend the root via its outermost tissues. We conclude that root angle is controlled by EGT1 in cereal crops employing an antigravitropic mechanism. (Pnas)
Brassinosteroid signaling restricts root lignification by antagonizing SHORT-ROOT function in Arabidopsis
Cell wall lignification is a key step in forming functional endodermis and protoxylem (PX) in plant roots. Lignified casparian strips (CS) in endodermis and tracheary elements of PX are essential for selective absorption and transport of water and nutrients. Although multiple key regulators of CS and PX have been identified, the spatial information that drives the developmental shift to root lignification remains unknown. Here, we found that brassinosteroid (BR) signaling plays a key role in inhibiting root lignification in the root elongation zone. The inhibitory activity of BR signaling occurs partially through the direct binding of BRASSINAZOLE-RESISTANT 1 (BZR1) to SHORT-ROOT (SHR), repressing the SHR-mediated activation of downstream genes that are involved in root lignification. Upon entering the mature root zone, BR signaling declines rapidly, which releases SHR activity and initiates root lignification. Our results provide a mechanistic view of the developmental transition to cell wall lignification in Arabidopsis thaliana roots.(Plant physiology)
Sulfate-TOR signaling controls transcriptional reprogramming for shoot apex activation
Plants play a primary role for the global sulfur cycle in the earth ecosystems by reduction of inorganic sulfate from the soil to organic sulfur-containing compounds. How plants sense and transduce the sulfate availability to mediate their growth remains largely unclear. The target of rapamycin (TOR) kinase is an evolutionarily conserved master regulator of nutrient sensing and metabolic signaling to control cell proliferation and growth in all eukaryotes. By tissue-specific Western blotting and RNA-sequencing analysis, we investigated sulfate-TOR signal pathway in regulating shoot apex development. Here, we report that inorganic sulfate exhibits high potency activating TOR and cell proliferation to promote true leaf development in Arabidopsis in a glucose-energy parallel pathway. Genetic and metabolite analyses suggest that this sulfate activation of TOR is independent from the sulfate-assimilation process and glucose-energy signaling. Significantly, tissue specific transcriptome analyses uncover previously unknown sulfate-orchestrating genes involved in DNA replication, cell proliferation and various secondary metabolism pathways, which largely depends on TOR signaling. Systematic comparison between the sulfate- and glucose-TOR controlled transcriptome further reveals that TOR kinase, as the central growth integrator, responds to different nutrient signals to control both shared and unique transcriptome networks, therefore, precisely modulates plant proliferation, growth and stress responses.(New Phytologist)
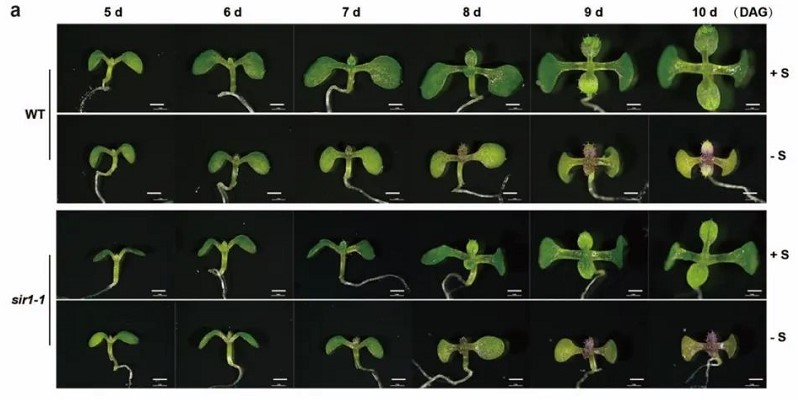

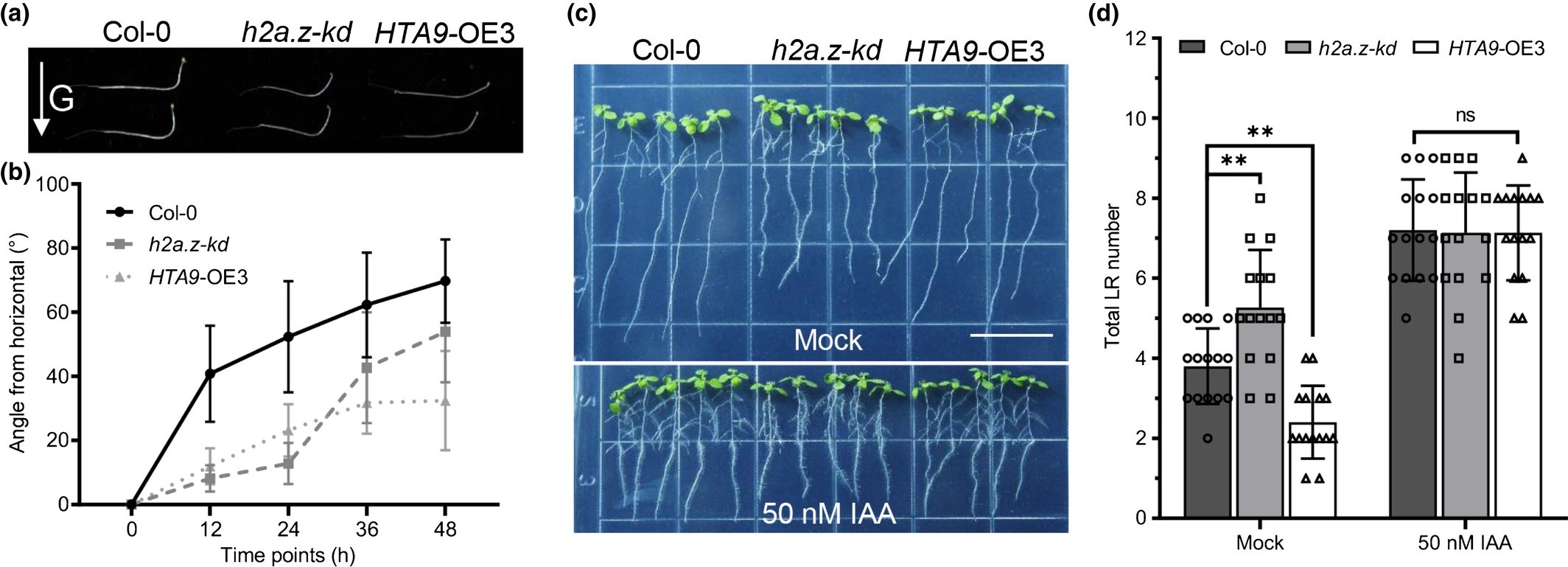
Feedback regulation of auxin signaling through the transcription of H2A.Z and deposition of H2A.Z to SMALL AUXIN UP RNAs in Arabidopsis
Auxin is a critical phytohormone that is involved in the regulation of most plant growth and developmental responses. In particular, epigenetic mechanisms, like histone modifications and DNA methylation, were reported to affect auxin biosynthesis and transport. However, the involvement of other epigenetic factors, such as histone variant H2A.Z, in the auxin-related developmental regulation remains unclear. We report that the histone variant H2A.Z knockdown mutant in Arabidopsis Col-0 ecotype, h2a.z-kd, has more lateral roots and weak gravitational responses related to auxin-regulated growth performances. Further study revealed that auxin promotes the eviction of H2A.Z from the auxin-responsive genes SMALL AUXIN-UP RNAs (SAURs) to activate their transcriptions. We found that IAA promotes the transcription of H2A.Z genes through HOMEOBOX PROTEIN 22/25 (AtHB22/25) transcription factors which work as downstream targets of ARF7/19 in auxin signaling. Double mutant of hb22 hb25 showed similar lateral root and gravitropism phenotypes to h2a.z-kd. Our results shed light on a reciprocal regulation hub through INOSITOL AUXOTROPHY 80-mediated H2A.Z eviction and ARF7/19-HB22/25-mediated H2A.Z transcription to modulate the activation of SAURs and plant growth in Arabidopsis.(New Phytologist)